Hiding in plain sight
The power in plant genomes isn’t just from genes – it’s also the ancient DNA switches we’re finally able to find
Professor Madelaine Bartlett takes us behind the scenes of a “needle in a haystack” search that uncovered ancient DNA sequences controlling plant genes – elements once thought not to exist.
Co-leader of the research that was published recently in Science, she explains how the international collaboration traced thousands of conserved regulatory elements back 300 million years, exposing fundamental principles of plant genome evolution and opening the door to more precise engineering of crop traits.

A deeply conserved DNA regulatory region shared across grasses helped shape maize domestication when a transposon (jumping gene) called “hopscotch” inserted itself near the TEOSINTE BRANCHED1 gene, it boosted its activity, suppressing branching and producing maize’s single, sturdy stem.
Cracking a long-standing challenge in plant genomics
When we think about genetics, we often think only about protein-encoding genes, or coding DNA.
However, once dismissed as ‘junk’ DNA, much of the action happens in the non-coding DNA.
In humans, 98% of our DNA is non-coding. In maize it’s 97%.
Indeed, the power in shaping plant traits, like increasing yields in crops, doesn’t lie in changing the proteins themselves, but in changing the DNA switches in non-coding DNA that control where, when and how strongly to turn on gene expression. These DNA switches are small segments of non-coding DNA called cis-regulatory elements.
Variation in non-coding DNA has long been recognised as a powerful evolutionary force. Back in 1975, Mary‑Claire King published a landmark paper showing that humans and chimpanzees share almost identical protein‑coding genes – about 99%.
She concluded that the big differences between our species arise not from the genes themselves, but from the regulatory sequences controlling them.
Since then, it’s become clear that regulatory evolution is also critical in plants. If we could identify cis-regulatory elements in genomes, we could understand how different species evolve, and also shape plant traits to meet human needs.
In animals, many cis-regulatory elements are maintained across evolutionary time as conserved non-coding sequences (CNSs).
Only a few ancient conserved non-coding sequences were identified in plant genomes. As a result, the consensus was that CNSs were rare in plants, and evolutionarily young. We weren’t convinced.
Plant genomes are huge and constantly reshuffling during evolution, so identifying CNSs was like trying to find a needle in a haystack. Maybe ancient CNSs were just hard to find in plant genomes.
The missing manual of plant evolution
In collaboration with Idan Efroni (The Hebrew University of Jerusalem) and Zachary Lippman (Cold Spring Harbor), we launched The Conservatory Project to see if we could locate these elusive CNSs.
We all research plant development – and were seeing hints that the regulation of developmental genes might be conserved between species.
Using high quality plant genomic data generated by the global plant research community over the past few years, we designed a new algorithm, Conservatory, to search for CNSs in genomes from 284 species across 72 plant families, spanning eudicots, monocots, gymnosperms, and algae.
Through the mammoth efforts of Kirk R. Amundson from University of Massachusetts Amherst and Anat Hendelman from Cold Spring Harbor Laboratory, we uncovered more than two million of these ancient regulatory sequences, some pre-dating the emergence of land plants 300 million years ago.
Our work, published in Science, finally revealed these regulatory sequences that were hiding in plain sight.
Many of these ancient sequences sit near genes involved in development – the genes that determine how plants grow, branch, flower, and form seeds.

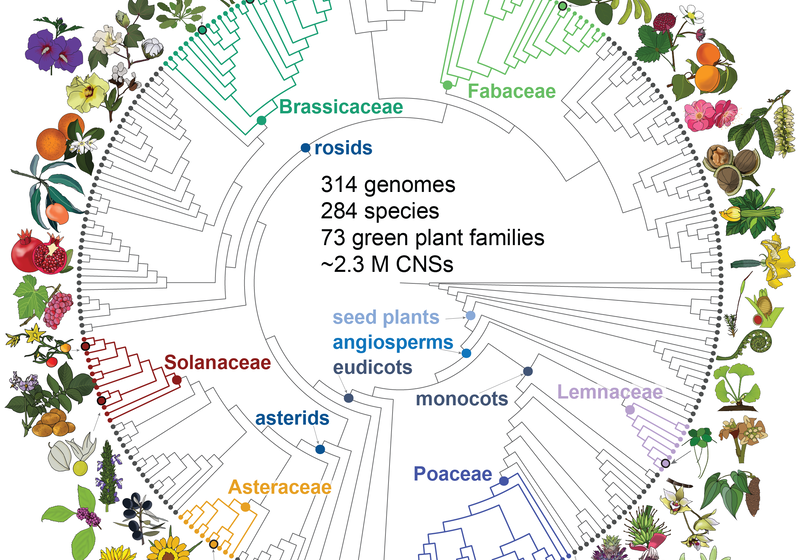
Phylogeny of the 284 species of plants included in Conservatory data set. Conservatory uncovered ~2.3 million conserved non-coding DNA sequences across 284 plant species from 72 families including eudicots, monocots, gymnosperms, and algae. Illustrations by Professor Madelaine Bartlett.
The grasses
We discovered in the Conservatory dataset a regulatory region that contains deeply conserved DNA switches that are shared across grasses.
Maize was domesticated from teosinte, a wild plant that has multiple stems.
During the domestication of maize, agronomists in Mexico selected plants with fewer branches, which eventually gave rise to maize’s iconic single sturdy stem.
We’ve long known this transformation was caused by a “hopscotch” transposon that jumped into the regulatory region near a gene called TEOSINTE BRANCHED1, boosting its expression and suppressing branching.
The hopscotch transposon just happened to land right on top of a regulatory region that had been shaping grass development for tens of millions of years.
For the first time, we can now identify these and other regions in grasses and other plant species.

Some plant lineages, including grasses (Poaceae) experienced particularly dramatic regulatory divergence early in their evolution. This dramatic regulatory rewiring may underpin the evolution of plant form, an area ripe for future discovery.
A roadmap for the next generation of crop engineering
Now that we know where many of these conserved regulatory sequences are, we have a roadmap for much more precise crop editing.
This is the subtle genetic toolkit that evolution has been using for hundreds of millions of years.
For my lab, and others, this dataset is a treasure trove.
We now have thousands of regulatory elements to explore, both to understand plant evolution, and to manipulate in agriculture.
We also haven’t found all the CNSs yet, but now have the tools to look.
Why these ancient ‘switches’ matter now
The ability to engineer crop traits with speed and precision is crucial as agriculture grapples with the triple threat of climate change, increasing disease, and rising food security demands.
Editing coding sequences is a heavy-handed approach. If you knock a gene out entirely, you often get drastic changes that are too abnormal for agricultural use.
What plant breeders want is the ability to ‘fine-tune’ traits. That’s the job of cis-regulatory elements.
For example, the CLAVATA3 gene in tomatoes plays a crucial role in regulating fruit size. If you mutate the CLAVATA3 gene itself you get big ugly misshapen tomatoes, but if you mutate the regulatory sequences, you get something more intermediate and useful. CLAVATA3 genes act similarly in maize.
Mutations in non-coding, regulatory DNA nudge a gene’s expression and function, causing, for example, a fruit to be slightly larger.
These subtle shifts are often exactly what agriculture needs.
Once dismissed as ‘junk’, identifying these ancient non-coding DNA sequences will be key for the future of crop trait editing.
Find out more
For more on The Conservatory Project, read our news article and the full research paper.