Locke Group

Professor James Locke
Associate Director of the Sainsbury Laboratory
Professor of Quantitative Plant Development
Research Group Leader
Email: james.locke@slcu.cam.ac.uk
Dynamic gene regulation from single cells to plant phenotypes
The Locke Group aims to understand how different types of gene expression dynamics are generated in plants and microbes and what their consequences are for growth, development and survival. Many genes are not simply "off" or "on". Even under constant external conditions, their expression levels change over time. For example, circadian clock genes oscillate with a period of about 24 hours, while other regulators show irregular pulses of activity that differ between genetically identical cells. These different types of dynamics are controlled by gene networks and influenced by stochasticity that arises from low molecule numbers. We want to understand the design principles of these circuits and how they operate in this noisy context.
To move from cataloguing gene networks to predicting and controlling plant phenotypes, we need to understand these dynamics. Static measurements tell us which genes are present and how active they are on average, but they miss how information is carried in temporal patterns, how variability between cells is generated or buffered, and why genetically identical plants can behave differently in the same conditions. By uncovering the design principles of noisy and dynamic gene regulation, we aim to develop a mechanistic understanding of how molecular circuits generate phenotypes in plants and microbes, and how similar principles are reused across systems. In the longer term, this understanding may also inform strategies to engineer plants that develop more uniformly yet remain resilient to environmental stress.
Key questions
We are currently focused on three overarching questions:
- What types of dynamic gene expression behaviours do cells use, and how are they encoded in circuit architecture? (For example, oscillations, stochastic pulses, bistable switches and slow fluctuations.)
- How are different dynamic pathways within a cell coupled to each other and to environmental signals? (For example, how clocks, stress pathways and developmental regulators interact.)
- How do single-cell dynamics, cell-to-cell variability and spatial coupling influence developmental decisions and stress responses at the whole-organism level? (For example, growth patterns and survival under stress.)
Our approach and model systems
We combine single-cell time-lapse microscopy, quantitative image analysis, single-plant and single-nucleus transcriptomics, stochastic and mechanistic modelling, and synthetic biology.
We focus on two complementary model systems:
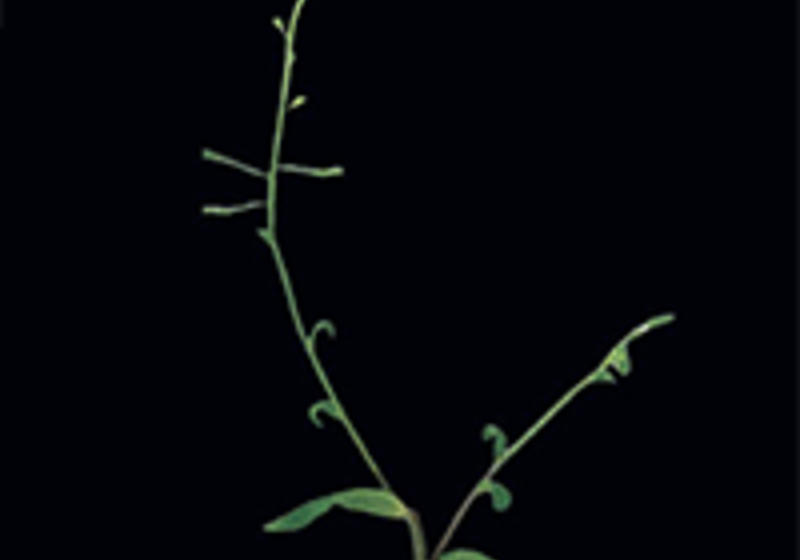
- Arabidopsis thaliana for multicellular plant development.
- Cyanobacteria and other microbes (including Bacillus subtilis) as simpler platforms to dissect and engineer noisy gene circuits.
Project 1 – Design principles of noisy gene circuits
Many of our microbial and synthetic biology projects ask how particular circuit motifs, especially mixed positive and negative feedback loops, feed-forward motifs and coupled oscillators, give rise to rich single-cell behaviours such as stochastic pulsing, bistability and oscillations.
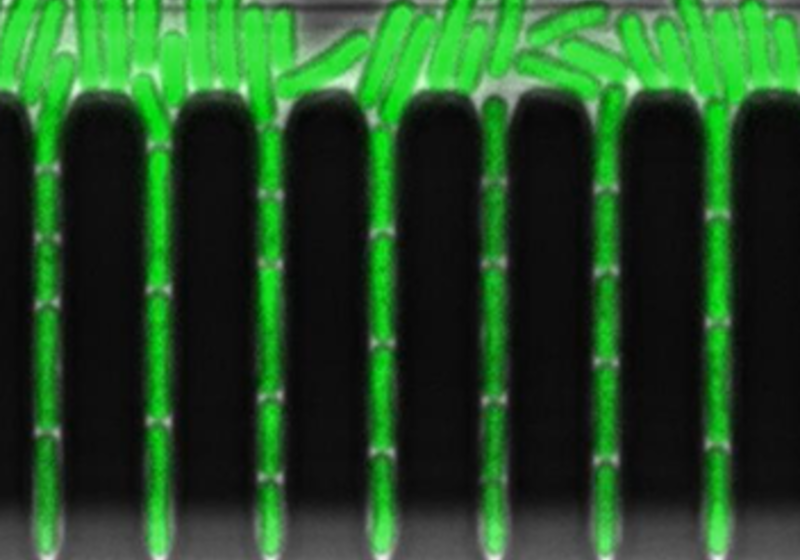
We use microfluidics and single-cell reporters to measure dynamics in individual bacteria, and stochastic modelling to map how behaviours change with circuit parameters. These studies provide design rules that help us interpret, and eventually re-engineer, gene circuits in more complex plant systems.
Project 2 – Coupling of dynamic gene networks
We study how dynamic gene networks, such as circadian clocks and stress-response pathways, are coupled to each other, to environmental and developmental signals, and between neighbouring cells.
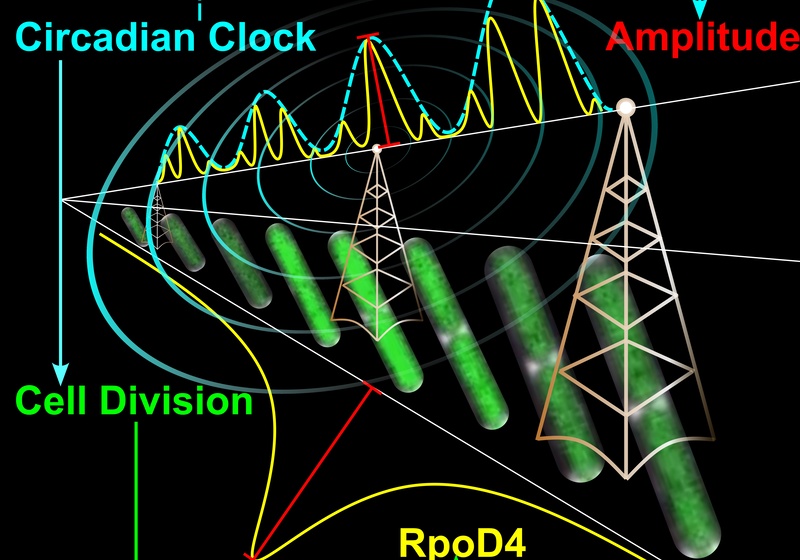
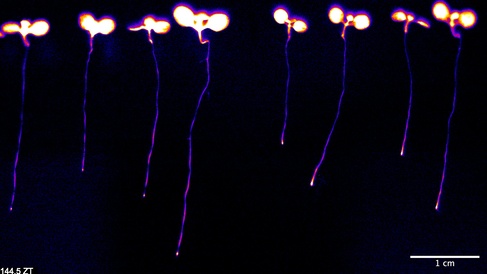
In cyanobacteria, we investigate how clock circuits couple to downstream targets and how dynamic strategies can expand the range of behaviours supported by a single clock. In plants, we use high-resolution luciferase imaging to map how clocks differ between organs and how local cell-to-cell coupling generates spatial waves that coordinate timing across the plant. These studies show how coupling within and between cells shapes the temporal and spatial organisation of physiology.
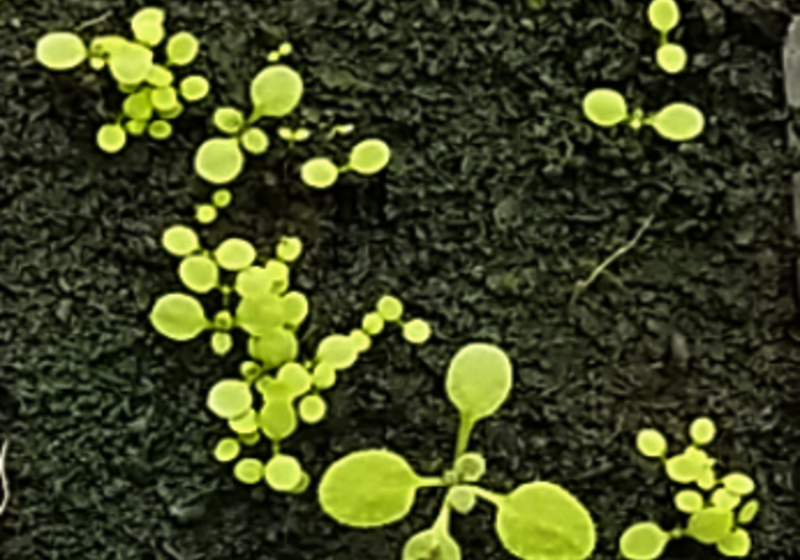
Research has found clocks in plant seedlings can self-organise without a master, collecting external signals such as light and temperature, and then communicating this information with their neighbours.
Project 3 – Dynamic regulation, variability and decision-making in plants
In plants, we investigate how noisy and dynamic gene expression influences developmental decisions and phenotypic variability, using Arabidopsis as a model.
Join or collaborate with the Locke Group
We welcome enquiries from prospective PhD students, postdocs, visitors and collaborators who are excited about single-cell dynamics, stochastic modelling or synthetic gene circuits in plants and microbes.
If you are interested in joining or collaborating with the group, please email james.locke@slcu.cam.ac.uk with a short description of your background and research interests.
Widespread inter‐individual gene expression variability in Arabidopsis thaliana
Coordinated circadian timing through the integration of local inputs in Arabidopsis thaliana
Cell size control driven by the circadian clock and environment in cyanobacteria
Nitrate modulates stem cell dynamics in Arabidopsis shoot meristems through cytokinins
View a full list of publications on Google Scholar
Selected publications
Ho-Wei Wu, James CW Locke (2026) Spatial Regulation of the Plant Circadian Clock, Annual Review of Plant Biology
Abhishek Upadhyay, Jamila Rowland-Chandler, Julia Stewart-Wood, Gabriela Pingarron-Cardenas, Isao T Tokuda, Alex Webb, James CW Locke (2026) Data-driven mathematical modelling explains altered timing of EARLY FLOWERING 3 in the wheat circadian oscillator, J R Soc Interface.
Poonam Bheda, Jingyi Hou, Ruedi Aebersold, Uri Alon, Joel S Bader, Lee Bardwell, Edison T Liu, James CW Locke, Matthias Mann, Andrew J Millar, Felix Naef, Yitzhak Pilpel, Ron Shamir, Dennis Vitkup (2025) Molecular Systems Biology at 20: reflecting on the past, envisioning the future, Molecular Systems Biology.
Z Nahas, AJ Bridgen, TE Loman, J Dillon, K Abley, DL Cano-Ramirez, F Ticchiarelli, M Seale, JCW Locke^, HMO Leyser^, A BRC1-modulated switch in auxin efflux accounts for the competition between Arabidopsis axillary buds, PLOS Biology, 2025; 23(9):e3003395. Research summary
A Eremina, C Schwall, T Saez, L Witting, D Kohlheyer, BMC Martins^, P Thomas^, JCW Locke^, Environmental and molecular noise buffering by the cyanobacterial clock in individual cells, Nature Communications, 2025; 16:3566. Research summary
C Ye, CN Micklem, T Saez, AK Das, BMC Martins^, JCW Locke^, The cyanobacterial circadian clock couples to pulsatile processes using pulse amplitude modulation, Current Biology, 2024; 34(24):5796–5803.e6. Research summary
M Greenwood, IT Tokuda, JCW Locke^, A spatial model of the plant circadian clock reveals design principles for coordinated timing, Molecular Systems Biology, 2022; 18(3):e10140.
CP Schwall, TE Loman, BMC Martins, S Cortijo, C Villava, V Kusmartsev, T Livesey, T Saez, JCW Locke, Tunable phenotypic variability through an autoregulatory alternative sigma factor circuit, Molecular Systems Biology, 2021; 17:e9832.
K Abley, P Formosa-Jordan, H Tavares, EYT Chan, M Afsharinafar, O Leyser^, JCW Locke^, An ABA-GA bistable switch can account for natural variation in the variability of Arabidopsis seed germination time , eLife, 2021; 10:e59485. Research summary
E Nadezhdin, N Murphy, N Dalchau, A Phillips, JCW Locke, Stochastic pulsing of gene expression enables the generation of spatial patterns in Bacillus subtilis biofilms,Nature communications, 2020; 11 (1), 1-12. Research summary
M Greenwood, M Domijan, PD Gould, AJW Hall, JCW Locke, Coordinated circadian timing through the integration of local inputs in Arabidopsis thaliana,PLoS biology, 2019; 17 (8), e3000407. Research summary and feature in The CoversationPlants can tell time even without a brain - here's how
S Cortijo, Z Aydin, S Ahnert, JCW Locke, Widespread inter-individual gene expression variability in Arabidopsis thaliana , Molecular systems biology, 2019; 15 (1), e8591. Research summary
O Patange, C Schwall, M Jones, C Villava, DA Griffith, A Phillips, JCW Locke, Escherichia coli can survive stress by noisy growth modulation,Nature communications, 2018; 9 (1), 1-11
BMC Martins, AK Tooke, P Thomas, JCW Locke, Cell size control driven by the circadian clock and environment in cyanobacteria,PNAS, 2018; (48), E11415-E11424
P Gould*, M Domijan*, M Greenwood, IT Tokuda, H Rees, L Kozma-Bognar, AJW Hall^, JCW Locke^, Coordination of robust single cell rhythms in the Arabidopsis circadian clock via spatial waves of gene expression,eLife, 2018; 7, e31700
Selected reviews
K Abley, R Goswami, JCW Locke, Bet-hedging and variability in plant development: seed germination and beyond, Philosophical Transactions of the Royal Society B, 2024; 379(1900):20230048
CN Micklem, JCW Locke, Cut the noise or couple up: Coordinating circadian and synthetic clocks, iScience, 2021; 24 (9), 103051
S Cortijo, JCW Locke, Does gene expression noise play a functional role in plants?Trends in Plant Science, 2020; 25 (10), 1041-1051
M Greenwood, JCW Locke, The circadian clock coordinates plant development through specificity at the tissue and cellular level,Current opinion in plant biology, 2020; 53, 65-72
*Joint first authors
^Joint corresponding authors
Abhishek Upadhyay (Research Associate) - next role as Senior Researcher, Institute for Advanced Study (IAS), Kyoto University, Japan
Enrico Sandro Colizzi (Research Associate) - next role as Tenured Scientist, INRIA, Lyon
Sasha Eremina (PhD Student) - next role as Entrepreneur in Residence, The Sainsbury Laboratory - Ventures (Norwich)
Torkel Loman (PhD Student) - next role as Postdoc, MIT
Mirela Domijan (Research Associate) - next role as Lecturer, University of Liverpool
Douglas Griffith (Research Assistant) - next role as Senior Researcher, LMU Munich
Casandra Villava (Research Assistant) - next role as Research Technician, EMBL Barcelona
Benoit Landrein (Research Associate) - next role as CNRS Independent Researcher
Niall Murphy (Research Associate) - next role at equal1.labs
Om Patange (PhD student) - next role as Postdoc, Harvard Medical School
Bruno Martins (Research Associate) - next role as Assistant Professor, University of Warwick
Sandra Cortijo (Research Associate) - next role as CNRS Independent Researcher
Pau Formosa-Jordan (Herchel Smith Post Doctoral Research Fellow) - next role as Group Leader, Max Planck Institute for Plant Breeding Research
Mark Greenwood (Research Associate) - next role as Postdoc, MIT
Research insights
Discover the story behind the research, from ideas to experiments and insights