Robinson Group

Research Group Interests
The Robinson Group uses a combination of novel biophysical tools, genetic manipulation and mathematical modelling to investigate how plant development (cell division and cell expansion) is controlled.
In plants the relationship between cell division and expansion is not clear. This is due to the rigid cell wall that surrounds the cells. Cell expansion requires the wall to extend and, therefore, depends upon how extensible it is and the forces that are acting on it, such as those due to turgor pressure. Cell expansion increases the size of the cell with little increase in biomass. On the other hand cells divide by adding more cell walls, increasing biomass with little increase in cell volume. Understanding this relationship may therefore enable us to improve the balance between plant size and biomass. This can be critical to optimise plants for biofuels, for which we want to maximise biomass for a given size.
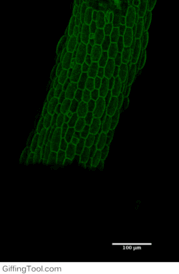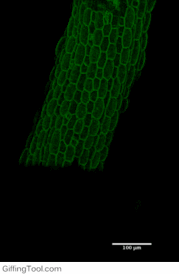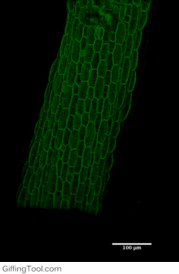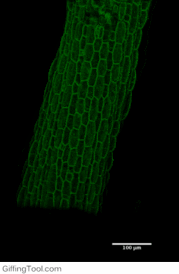
Cell wall impact on tissues
How cell division impacts cell expansion has been a subject of controversy for decades and has not yet been solved by either molecular or genetic approaches. The main reason for this is due to a lack of suitable measurement methods. Group Leader, Sarah Robinson developed an automated confocal micro-extensometer (ACME) that enables mechanical measurements to be made in 3D at the scale of the tissue but with cellular resolution. The Robinson Group is using ACME to determine the impact of new cell walls on the properties of the tissue.
Mechanical stress feedback
Feedback regulation by mechanical stress on development has recently re-emerged as an important mechanism for plant development. The Robinson Group are also assessing the relationship between the mechanical properties and cell division using ACME. This involves applying precise mechanical stress then inducing cell divisions and observing their orientation.
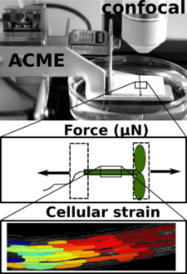
ACME robotic system
The Robinson Research Group utilises a robotic system developed by Dr Sarah Robinson, called ACME (automated confocal micro-extensometer) to measure the mechanical properties of plants in vivo (Robinson 2017 et al). ACME functions like a traditional extensometer, but is miniaturised, mounted on the stage of a confocal microscope, and fully automated.
ACME computes the mechanical properties of samples at the cellular level based on changes in features of the tissue itself (tracked in time-lapse z-stack images) during application of a known force or deformation.
There are a wide range of quantitative biophysical applications that the technology can be used for, including for analysis of the mechanical properties of developing plant organs.
Key Publications
*Bou Daher, F., Serra, L., Carter, R., Jönsson, H., Robinson, S., Meyerowitz, E.M. and Gray, W.M. (2024) Xyloglucan deficiency leads to a reduction in turgor pressure and changes in cell wall properties, affecting early seedling establishment, Current Biology, 34(10), 2094-2106.e6 doi: 10.1016/j.cub.2024.04.016
*Sénéchal, F., Robinson, S., Van Schaik, E., Trévisan, M., Saxena, P., Reinhardt, D. and Frankhauser, C. (2024) Pectin methylesterification state and cell wall mechanical properties contribute to neighbor
proximity-induced hypocotyl growth in Arabidopsis, Plant Direct, 8(4), e584 doi: 10.1002/pld3.584
Alessandra Bonfanti, Euan Thomas Smithers, Matthieu Bourdon, Alex Guyon, Philip Carella, Ross Carter, Raymond Wightman, Sebastian Schornack, Henrik Jönsson, Sarah Robinson (2023) Stiffness transitions in new walls post-cell division differ between Marchantia polymorpha gemmae and Arabidopsis thaliana leaves. PNAS DOI: 10.1073/pnas.2302985120
Robinson, S. (2021), Mechanobiology of cell division in plant growth. New Phytol, 231: 559-564. https://doi.org/10.1111/nph.17369
Chiara A. Airoldi, Carlos A. Lugo, Raymond Wightman, Beverley J. Glover, Sarah Robinson* (2021) Mechanical buckling can pattern the light-diffracting cuticle of Hibiscus trionum. Cell Reports DOI: 10.1016/j.celrep.2021.109715 (*co-corresponding)
Serra L, Robinson S. (2020) Plant cell divisions: variations from the shortest symmetric path. Biochem Soc Trans. DOI:10.1042/BST20200529 (corresponding author).
Yarahmadov* T, Robinson* S, Hanemian* M, Pulver V, Kuhlemeier C. (2020) Identification of transcription factors controlling floral morphology in wild Petunia species with contrasting pollination syndromes. The Plant Journal.DOI:10.1111/tpj.14962 (co-first authors)
Bonfanti A, and Robinson S. (2020). Studying cell wall mechanics using an automated confocal micro-extensometer. Methods in Cell Biology, Academic Press. DOI: 10.1016/bs.mcb.2020.04.001
Blösch R, Plaza-Wüthrich S, Barbier de Reuille P, Weichert A, Routier-Kierzkowska AL, Cannarozzi G, Robinson S*, Tadele Z*. (2020). Panicle angle is an important factor in tef lodging tolerance. Front Plant Sci.10.3389/fpls.2020.00061 (co-corresponding author).
Robinson, S. and Durand-Smet, P. (2020) Combining tensile testing and microscopy to address a diverse range of questions. Journal of Microscopy.doi:10.1111/jmi.12863
Robinson, S. and Kuhlemeier, C. (2018) Global compression reorients cortical microtubules in Arabidopsis hypocotyl epidermis and promotes growth.Current Biology (2018) 28:1794–1802. https://doi.org/10.1016/j.cub.2018.04.028
- A quantitative assessment of microtubule responses to in-plane application of mechanic forces was conducted and compared to the stresses predicted from a finite element model.
Melnyk, C.W., Gabel, A., Hardcastle, T.J., Robinson, S., Miyashima, S., Grosse, I., Meyerowitz, E.M. (2018) Transcriptome dynamics at the Arabidopsis graft junction reveal an inter-tissue recognition mechanism that activates vascular regeneration.PNAS 115:2447-2456. https://doi.org/10.1073/pnas.1718263115
Samantha Fox, Paul Southam, Florent Pantin, Richard Kennaway, Sarah Robinson, Giulia Castorina, Yara E Sánchez-Corrales, Robert Sablowski, Jordi Chan, Verônica Grieneisen, Athanasius FM Marée, J Andrew Bangham, Enrico Coen. (2018). Spatiotemporal coordination of cell division and growth during organ morphogenesis. PLoS biology 16 (11), e2005952
Robinson, S., Huflejt, M., Barbier de Reuille, P., Braybrook, S.A, Schorderet, M., Reinhardt, D., Kuhlemeier, C. (2017) An automated confocal micro-extensometer enables in vivo quantification of mechanical properties with cellular resolution.The Plant Cell 29:2959-2973. https://doi.org/10.1105/tpc.17.00753
- We expand the possibility of making quantitative mechanical measurements in the plane of the tissue while live imaging with a confocal microscope.
Barbier de Reuille, P., Routier-Kierzkowska, A.-L., Kierzkowski, D., Bassel, G. W., Schüpbach, T., Tauriello, G., Bajpai, N., Strauss, S., Weber, A., Kiss, A., Burian, A., Hofhuis, H., Sapala, A., Lipowczan, M., Heimlicher, M. B., Robinson, S., Bayer, E. M., Basler, K., Koumoutsakos, P., Roeder, A. H., Aegerter-Wilmsen, T., Nakayama, N., Tsiantis, M., Hay, A., Kwiatkowska, D., Xenarios, I., Kuhlemeier, C. & Smith, R. S. (2015) MorphoGraphX: A platform for quantifying morphogenesis in 4D.elife 4:e05864. https://elifesciences.org/articles/05864. https://doi.org/10.7554/eLife.05864.
- A popular software to aid quantitative image analysis.
Barbier de Reuille, P., Robinson, S., Smith, R. S. (2014) Quantifying Cell Shape and Gene Expression in the Shoot Apical Meristem Using MorphoGraphX. Humana Press, 121-134. https://doi.org/10.1007/978-1-62703-643-6_10
Robinson, S., Burian, A., Couturier, E., Landrein, B., Louveaux, M., Neumann, E., Peaucelle, A., Weber, A., Nakayama N. (2013) Mechanical control of morphogenesis at the shoot apex.Journal of Experimental Botany. 64:4729:4744. # co-corresponding author. https://doi.org/10.1093/jxb/ert199
Nakayama, N., Smith, R., Mandel, T., Robinson, S., Kimura, S., Boudaoud, A., Kuhlemeier, C. (2012) Mechanical regulation of auxin-mediated growth.Current Biology 22:1468-1476. https://doi.org/10.1016/j.cub.2012.06.050
- Evidence that the cellular localisation of the auxin transporter PIN1 is sensitive to mechanical stress.
Kuchen, E., Fox, S., Barbier de Reuille, P., Kennaway, R., Bensmihen, S., Avondo, J., Calder, G., Southam, P., Robinson, S., Bangham, A., Coen, E. (2012) Generation of leaf shape through early patterns of growth and tissue polarity. Science 335:1092-1096. https://doi.org/10.1126/science.1214678
*Robinson, S., Barbier de Reuille, P., Chan, J., Bergmann, D.C., Prusinkiewicz, P., and Coen, E.S. (2011) Generation of spatial patterns through cell polarity switching.Science 333:1436-1440. http://science.sciencemag.org/content/333/6048/1436
- by combining modelling with time-lapse imaging we are able to predict the pattern of cell division and inherited cell identity in a growing leaf.
Pre-Print Publications
Robinson, S., Huflejt, M., Barbier de Reuille, P., Braybrook, S.A, Schorderet, M., Reinhardt, D., Kuhlemeier, C. (2017) An automated confocal micro-extensometer enables in vivo quantification of mechanical properties with cellular resolution. bioRxiv 183533; https://doi.org/10.1101/183533
Melnyk, C.W., Gabel, A., Hardcastle, T.J., Robinson, S., Miyashima, S., Grosse, I., Meyerowitz, E.M. (2017) Transcriptome dynamics at the Arabidopsis graft junction reveal an inter-tissue recognition mechanism that activates vascular regeneration. bioRxiv 198598; https://doi.org/10.1101/198598https://www.biorxiv.org/content/10.1101/198598v2
Samantha Fox, Paul Southam, Florent Pantin, Richard Kennaway, Sarah Robinson, Giulia Castorina, Yara E Sánchez-Corrales, Robert Sablowski, Jordi Chan, Verônica Grieneisen, Athanasius FM Marée, J Andrew Bangham, Enrico Coen. (2018). Spatiotemporal coordination of cell division and growth during organ morphogenesis. PLoS biology 16 (11), e2005952 Paulina Piepzyca 16.00 Normal 0 false false false EN-GB X-NONE X-NONE
