Jönsson Group

Professor Henrik Jönsson
Sainsbury Laboratory Director
Research Group Leader
Professor of Computational Morphodynamics
Sainsbury Laboratory | University of Cambridge
Email: Henrik.Jonsson@slcu.cam.ac.uk
Computational morphodynamics in plant development
Henrik Jönsson’s research group develops computational morphodynamics models at the cellular level to describe multicellular plant tissues such as the shoot apical meristem. The group works in close collaboration with experimental partners.
The models integrate gene regulatory networks, hormone transport and signalling, cell growth and division, and mechanical properties. The group continuously evaluates and refines model structure and parameters using new experimental data, primarily from live microscopy.
Integrating modelling and experimental data
The group combines live imaging, genetic and hormonal perturbations, and single-cell transcriptomics with theoretical and computational modelling. This approach supports a quantitative understanding of plant development across multiple scales.
The group integrates molecular and mechanical mechanisms to link sub-cellular processes with tissue-level development in plant shoot meristems and other tissues.
Shoot apical meristem development
The group studies the development and regulation of the shoot apical meristem (SAM), focusing on how new primordia form at the meristem periphery.
Auxin accumulates at sites where new organs form, supported by feedback mechanisms that regulate its transport. Mechanical stresses contribute to intercellular transport and influence fibre formation, helping regulate primordium growth and positioning.
The models incorporate molecular reactions, transport and signalling, physical stress and growth, and interactions between these processes.
The group also investigates how the SAM maintains stable cell differentiation patterns throughout plant life, despite continuous cell replacement during symplastic growth.
A key focus is the CLAVATA–WUSCHEL negative feedback network, which regulates gene expression and intercellular signalling. The group optimises and evaluates model parameters using large sets of perturbation experiments, with a focus on identifying parameter regions that explain network behaviour. Receptor cross-talk within this regulatory system is also studied.
Research approach
The group uses iterative cycles of modelling and experimental validation. Models are continuously tested against new data to improve accuracy and predictive power. Live microscopy data plays a central role in this process.
Future directions
The group aims to build integrated, multi-scale models of plant development that connect molecular regulation, mechanics, and tissue growth. This includes improved mapping of experimental data onto virtual tissues and the use of computational and AI-based methods to analyse large datasets and generate testable hypotheses.
The research also extends to the interaction between meristem, stem, and floral tissues, including vascular development, and explores de novo meristem formation from root tissues to understand self-organisation processes.
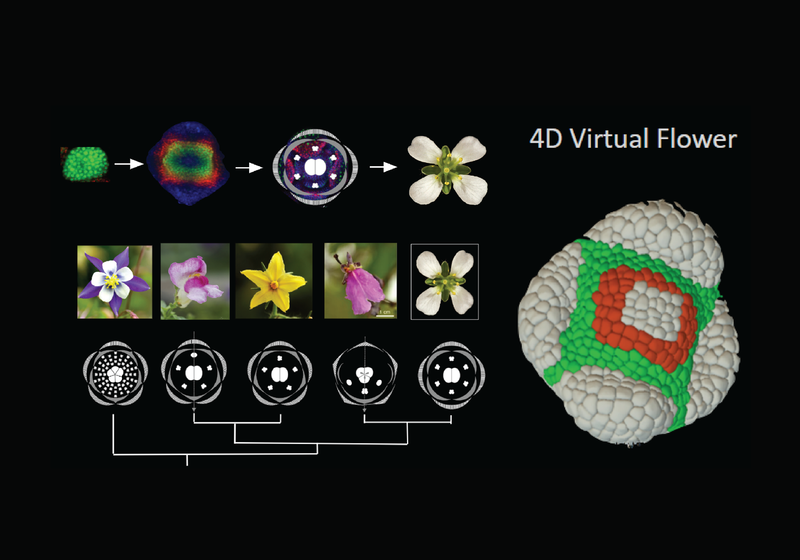
Building a virtual flower
ERC Synergy grant awarded to a collaboration including the Jönsson Group at the Sainsbury Laboratory, Europe and Australia to build a virtual flower and study symmetry breaking in multicellular organisms, addressing how complex biological forms are generated.
View full list of Jönsson Group publications on Google Scholar
Selected publications
CSLD-5-mediated cell wall remodelling regulates tissue mechanics and shoot meristem growth
Journal article | 2025 | Nature Communications
Lan, M., Zhu, Y., Peaucelle, A., Zhu, X., Liu, Y., Cao, X., Gurzadyan, A., Gao, G., Cai, W., Gruel, J., Haas, K., Jönsson, H.,
Hamant, O., Wightman, R., Meyerowitz, E. and Yang, W.
Differential growth in an emergent property of mechanochemical feedback mechanisms in curved plant organs
Journal article | 2024 | Developmental Cell
Walia, A., Carter, R., Wightman, R., Meyerowitz, E.M., *Jönsson, H. and *Jones, A.M.
Hibiscus bullseyes reveal mechanisms controlling petal proportions that influence plant-pollinator interactions
Journal article | 2024 | Science Advances
Riglet, L., Zardilis, A., Fairnie, A.L., Yeo, M.T.S., Jönsson, H. and *Moyroud, E.
Sugar signalling modulates SHOOT MERISTEMLESS expression and meristem function in Arabidopsis
Journal article | 2024 | PNAS
Lopes, F.L., Formosa-Jordan, P., Malivert, A., Margalha, L., Confraria, A., Feil, R., Lunn, J.E., *Jönsson, H., *Landrein, B.
and *Baena-González, E.
Mean-field theory approach to three-dimensional nematic phase transitions in microtubules
Review | 2023 | Physical Review E
Gibson, C., *Jönsson, H. and *Spelman, T.
Stiffness transitions in new walls post-cell division differ between Marchantia polymorpha gemmae and Arabidopsis thaliana leaves
Journal article | 2023 | PNAS
Bonfanti, A., Smithers, E.T., Bourdon, M., Guyon, A., Carella, P., Carter, R., Wightman, R., Schornack, S., Jönsson, H. and
*Robinson, S.
A 3D gene expression atlas of the floral meristem based on spatial reconstruction of single nucleus RNA sequencing data
Journal article | 2022 | Nature Communications
Neumann, M., Xu, X., Smaczniak, C., Schumacher, J., Yan, W., Blüthgen, N., Greb, T., Jönsson, H., Traas, J., Kaufmann, K.
and *Muino, J.
Molecular mechanism of cytokinin-activated cell division in Arabidopsis
Journal article | 2021 | Science
Yang, W., Cortijo, S., Korsbo, N., Roszak, P., Schiessl, K., Gurzadyan, A., Wightman, R., *Jönsson, H. and Meyerowitz,
E.M.
Tissue folding at the organ-meristem boundary results in nuclear compression and chromatin compaction
Journal article | 2021 | PNAS
Fal, K., Korsbo, N., Alonso-Serra, J., Teles, J., Liu, M., Refahi, Y., Chaboute, ME., *Jönsson, H. and Hamant, O
A multiscale analysis of early flower development in Arabidopsis provides an integrated view of molecular regulation and growth control
Journal article | 2021 | Developmental Cell
Refahi, Y., Zardilis, A., Michelin, G., Wightman, R., Leggio, B., Legrand, J., Faure, E., Vachez, L., Armezzani, A., Risson,
AE.,Zhao, F., Das, P., Prunet, N., Meyerowitz, E.M., *Jönsson, H. and Traas, J.
