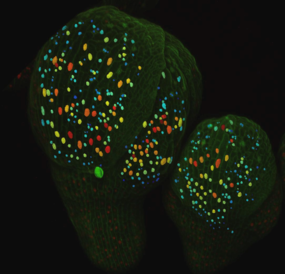
A key regulator has been discovered to determine cell size in flowers through random fluctuations. Counterintuitively, this randomness can lead to patterns. This finding helps us to understand how biological patterns are initiated, how shape and size are determined during growth, and may lead to important discoveries improving crop yields.
Scientists from the Locke and Jönsson group at the Sainsbury Laboratory, University of Cambridge collaborated with the Roeder group at Cornell University (USA), to investigate the formation of giant cells in sepals, the outer organ of a flower. The proportion of giant cells to small cells is of importance because it allows the sepal to have the right shape and perform its protective function in the flower.
By combining live imaging of single cells with advanced image processing methods and computational modeling, scientists found that ATML1, a protein important for determining cell fate, fluctuated over time in the sepal. Initial experiments where the levels of ATML1 were changed showed that the number of giant cells followed the amount of ATML1, while the computational model predicted that high levels of ATML1 could be decisive for a single cell to stop dividing and become a giant cell.
Being able to quantify and follow these fluctuations at the single cell level in the growing plant as it develops was crucial for correlating the expression dynamics of this key regulator with cell fate decisions, and provided a powerful experimental basis for the study. The computational tools developed at SLCU were essential for quantitatively understanding how the regulation was achieved. ‘The work is a nice example where a computational model is integral for understanding a biological process, a method that is standard in physics research’, says Pau Formosa-Jordan, a researcher at the SLCU and lead author of the study together with José Teles, and Heather Meyer from Cornell.
These results provide experimental evidence for a novel mechanism of cell fate decisions, where fluctuations lead to patterns, which could be of importance in other plant tissues. More broadly, due to their fundamental nature, the observations can also be of importance for other multicellular systems beyond plants. ‘Animal cells frequently experience processes of decision-making, and these decisions underlie fundamental aspects from embryonic development to tissue homeostasis in health and disease’ says José Teles.
Computational modelling played a fundamental role in connecting the observed fluctuations in ATML1 to patterns in cell size. This integrated research approach combining modelling, live imaging and detailed image quantification, has the potential to transform how life science research is performed and, in the long term, provide efficient tools for increasing yield in agriculture and biofuel production.
The finding is published in the scientific journal eLife.
Contact:
| James Locke | Henrik Jönsson |
| james.locke@slcu.cam.ac.uk | henrik.jonsson@slcu.cam.ac.uk |
| +44 1223 761110 | +44 1223 761128 |
