SLCU's François Nédélec has joined Andrea Musacchio, from the Max Planck Institute of Molecular Physiology in Dortmund, and Thomas Surrey, from the Centre for Genomic Regulation in Barcelona, to study one of the most fundamental processes in life – cell division. Their project has been awarded a prestigious ERC Synergy Grant, valued at 11 million Euros, to reconstitute cell division in vitro and in silico
When a cell divides, it must go through a radical and dramatic transformation before it splits. Chromosomes are duplicated, rearranged and aligned in a very particular manner before they are pulled into the two daughter cells. Despite this mechanism being at the heart of biology, it is still poorly understood. The collaborators have set the extraordinarily ambitious goal of unravelling this interplay by re-engineering it in vitro and by modelling it in silico.
Pooling their resources, they hope to reconstitute and model cell division on an entirely new level. Their project has been awarded one of the rare Synergy Grants from the European Research Council. The partners in the consortium have already gained groundbreaking insight into the understanding of cell division in recent years, and their collaboration will uncover new fundamental insights of living system.

Cell division is regulated by networks with a large number of proteins. Just like the instruments in an orchestra, proteins in networks must work together in a concerted manner. The smallest error can lead to noticeable malfunctions, and in the case of cell division errors may cause developmental disorders or the development of cancer.
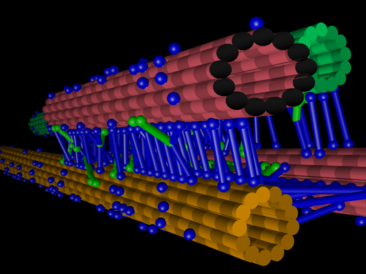
The larger the orchestra becomes, the more difficult it is to coordinate its players. This problem is particularly relevant when studying complex protein networks, such as the biochemical and mechanical networks driving cell division. In living cells, there are simply too many unknown players influencing this sophisticated process, making precise investigations of the role of individual players very arduous. The mitotic spindle is key to this as the microtubule-based bipolar structure that segregates the chromosomes in cell division.
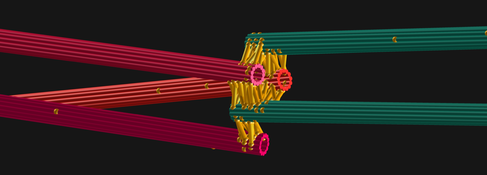
Research Group Leader at SLCU, Dr Nédélec says the collaboration's goal is to generate a quantitative understanding, grounded on the first principles of physics and chemistry, of how the cell cycle and the mitotic spindle achieve chromosome segregation.
"We have been modelling cell division for more than 20 years, studying molecular machines like the mitotic spindle and for a physicist, this is a wonderful playground," says Dr Nédélec.
"Just like a regular engine, the mitotic spindle consumes energy to perform mechanical work, but it is built on totally different principles. As a theoretician, I need collaborators who are experts in the biology and working together will give us deeper insights into the complexity of cell division. With this iconic example, we plan to illustrate how inanimate biochemical components are turned into living matter."
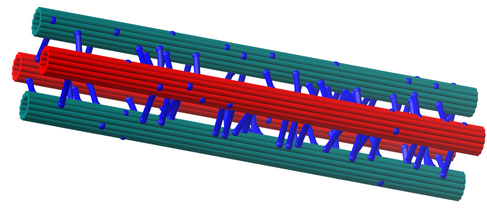
Reconstitution to an entirely new level
The biochemists Andrea Musacchio and Thomas Surrey follow an approach known as biochemical reconstitution, where the protein network of interest is reduced to a manageable but still functional set of players and analysed in a controllable environment. To do so, the reduced set is transferred out of the living system into a test tube and reconstituted - protein by protein. "Only by taking the system apart and simplifying it do we have a chance of understanding how cell division works. That is why we want to re-engineer it in the laboratory as accurately as possible and without compromising on complexity," explains Musacchio, principal investigator of the joint project.
Both scientists are leading experts in the field of in vitro reconstitution and dynamic analysis of complex biochemical systems. Over the past two decades, they have put a lot of work and experience into building up exceptional libraries of purified human proteins from four fundamental networks that are iconic of eukaryotic cell division, including the spindle assembly checkpoint, the metaphase spindle, the kinetochores and the central spindle.
"While we have studied the four networks separately in the past, we will combine our library and our experience and add further components to increase the complexity and significance of our reconstitutions and thus achieve an unprecedented depth of analysis," says Surrey.
The data obtained from this analysis will be interpreted using mathematical models and simulations. An expert in this field and the third partner in the consortium is François Nédélec. "The more reduced the studied network, the fewer unknowns are in the problem and the better mathematical models can be developed that take time and space into account. This is the key to a physically realistic simulation, in which, ultimately, individual parameters can be changed to predict the resulting effects on the entire system.”
"The idea underlying our project is to use a defined set of real proteins combined with computer-generated simulations to decipher how protein networks in the cell generate sophisticated biological functions. In our case, this is the correct transmission of genetic information during cell division, crucial for genetic inheritance, reproduction of life and development of any organism. Ideally, our synergistic approach will uncover general principles of how living systems work" says Andrea Musacchio, describing the project's grand vision.
About the ERC
The European Research Council, set up by the European Union in 2007, is the premier European funding organization for excellent frontier research. Every year, it selects and funds the most outstanding, creative researchers of any nationality and age to run projects based in Europe. The ERC has three grant schemes for individual principal investigators – Starting Grants, Consolidator Grants, and Advanced Grants – and Synergy Grants for small groups of excellent research teams with up to €14 million in funding for up to six years.
In the 2019 competition for these grants, 37 consortia were selected for funding. A total worth of €363 million, these special grants will enable groups of two to four top researchers to bring together complementary skills, knowledge, and resources in one research project. The recipients will be able to tackle some of the most complex research problems, often spanning multiple scientific disciplines. This funding is part of the EU's research and innovation program, Horizon 2020.
More information
